Microbial Ecology and Diversity
Understanding the role of microorganisms in ecological processes and the spatiotemporal responses of microbial biodiversity to global change.
Carbon and Nitrogen acquisition and cycling in heterotrophic and mixotrophic ciliates
George McManus is interested in microbial eukaryotes and their roles in marine food webs. He is focused mostly on ciliates, which are among the most abundant herbivores in the sea. These organisms often practice a kind of mixotrophy in which chloroplasts from ingested food are retained and used to provide an energy subsidy for growth. His lab has successfully cultivated a number of these fastidious organisms. Using local isolates, he uses isotopes of C and N to study their metabolism. He is also interested in the biogeography of ciliates and uses DNA sequencing methods to evaluate their distributions and diversity.
Composition and development of the characteristic Thermotoga maritima cell sheath or "toga"
Dr. Noll’s laboratory studies the physiology and molecular genetics of hyperthermophilic bacteria and archaea to shed light on their mechanisms of evolution, cellular structure, and potentials for biofuel generation. Strictly anaerobic, extremely thermophilic bacteria belonging to the group Thermotogales are the primary organisms under study. Our evolutionary studies focus on how the evolution of sugar transporters including their horizontal transfer among bacteria and archaea affects the binding properties of their proteins. Another research focus involves the unique cell envelope found in the Thermotogales, a structure with features perhaps reminiscent of cell barriers found in ancestors of bacteria and archaea. Finally a new effort to engineer thermophiles for use in biofuels generation is underway, an effort that seeks to address some of the major impediments to biomass utilization. Full genome sequences for several members of the Thermotogales have become available due to the efforts of our lab and this information is being used to guide these studies. Genomic and proteomic approaches are being used for these projects.
Environmental biotechnology
Research in the Li lab focuses on bioresource recovery, carbon sequestration, environmental sensor development, real-time water/soil quality monitoring, energy-positive wastewater treatment
Evolution and ecology of variation within species; evolutionary immunology; host-parasite co-evolution; speciation.
Research in Daniel Bolnick's lab seeks to understand how ecological interactions affect the evolution of within-species trait variation. Research in the lab touches on a wide variety of species interactions, and combines theoretical models, natural history, field and lab experiments, and meta-analyses. Currently evolution of vertebrate immunity to parasites is a major, but not exclusive, focus of the lab.
Exploration of the structure and function of the bivalve-associated microbiome
Evan Ward’s laboratory studies the bivalve-associated microbiome to understand its involvement in host physiological processes. Bivalves are important members of coastal ecosystems and have commercial value, yet their microbiomes remain largely unexplored. We are interested in the role of gut/digestive gland and gill microbiota in bivalve toxicology, specifically involving anthropogenic pollutants like microplastics, nanoparticles, and pesticides. We utilize traditional bivalve husbandry and toxicological methods along with high-throughput sequencing and culture-based microbial techniques to conduct this research.
Genomics, proteomics and transcriptomics of bacteria (actinomycetes) that infect higher plants and fix atmospheric nitrogen; diversity of fungi associated with surface ripened cheese
Benson's research interests include microbial biogeography, including the molecular and genomic interactions of bacteria with their environment, and the genomic evolution controlling distribution of microbes in several environments. The specific areas of interest include plant-microbe and insect-microbe microbiomes, and the distribution and diversity of microorganisms associated with cheese.
Identifying archaea in microbial communities
Methane-producing archaea (also known as methanogenic archaea or methanogens) are free-living organisms that typically live in diverse free-oxygen environments, but some species have been also found in host-associated microbiomes. The Microbial Ecophysiology Lab (Santiago-Martinez Lab) is also interested in analyzing the diversity and function of methanogens in diverse environments. Our goal is to identify archaea associated with complex microbial communities and their role in One Health (the health of humans, animals and ecosystems).
Microbial systems influenced by micro-scale habitat features
Leslie Shor’s research group studies microbial systems influenced by micro-scale habitat features. We have been developing microfluidic devices to replicate physical, chemical, and biological features of the rhizosphere microbial system. In collaboration with the Gage lab, we have been investigating the effects of protozoan predation on biofilm formation of soil microbes; of microbially-produced extracellular polymers on soil moisture retention; and on protist-facilitated dispersal of bacteria in microfluidic networks.
Multipartite Symbioses as Models to Understand the Diversity, Stability and Function of Natural Microbial Communities
Jonathan Klassen studies microbial community ecology, especially using the fungus-growing ant symbiosis as a model system to understand how microbial interaction networks evolve. He is particularly interested in using relatively simple multipartite symbioses as models to understand the evolutionary and ecological rules governing the stability and function of microbial communities more generally. The Klassen lab uses culture-dependent and -independent genomics coupled with comparative phenotyping and chemical biology to understand how the differing ecologies and population structures of microbes govern the stability and function of the communities to which they belong.
Role of gut microbiota in host defenses against parasites
Sarah Knutie is a disease ecologist and evolutionary biologist who studies the role of gut microbiota in host defenses against parasites. Specifically, she’s interested in whether and how the gut microbiota affects developmental immunity to micro- and macroparasites in birds, frogs, and fish. Notably, her lab works in the Galapagos Islands where she studies the effect of an emerging disease on Darwin’s finches and how urbanization can influence host defense mechanisms against the parasite.
Understanding how beneficial symbioses are established and maintained
The Nyholm laboratory explores beneficial host/microbe interactions, ultimately trying to understand the mechanisms by which bacterial symbionts and their animal hosts communicate and how the host’s innate immune system might influence these interactions. We are currently studying how these complex associations are established, regulated and maintained. We use the model association between the Hawaiian bobtail squid, Euprymna scolopes, and the bioluminescent bacterium Vibrio fischeri, to to understand mechanisms of host/symbiont specificity. We are also exploring a bacterial consortium that is found in a reproductive gland of female squid and is hypothesized to provide antimicrobial and/or antifouling compounds in order to protect the developing, externally laid embryos.
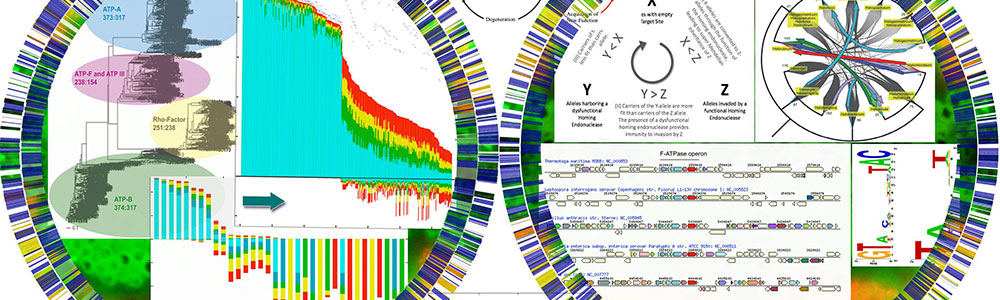
Understanding how genetic variation is distributed within and between haloarchaeal species and how molecular mechanisms for gene transfer work in Archaea
Papke lab Although we recognize names of microorganisms like Escherichia coli and Bacillus subtilis, our ability to classify strains into ³natural kinds² is rigorously tested by the observation that genetic variation is frequently shuttled across so-called species boundaries. Also, much of what we know about species comes from well-studied pathogenic bacteria, which are often classified by the disease that they cause (e.g., Bacillus anthracis or Neisseria gonorrhoeae). This is important to doctors and their patients but the Earth¹s biomass and diversity is comprised mainly of prokaryotes, the vast majority of which do not cause disease. Combined, these observations suggest that our understanding of species evolution is relatively shallow and that our current standards of classification are biased and difficult to apply to most microbes. The main goal of our research is to use evolution and ecology theory in combination with approaches like genomics, metagenomics and population genetics to investigate intra and inter species variation, gene flow and genetic relationships for non-pathogenic (e.g., environmental) prokaryotes. We concentrate on hypersaline adapted Archea (Haloarchaea) as model organisms because they live in island-like habits which aid in simplifying or sorting the evolutionary forces that effect their distributions, adaptations and variation.
Understanding spatiotemporal changes in multiple dimensions of microbial biodiversity in disturbance-mediated tropical systems
Willig lab: As part of the NSF-funded Luquillo Mountains Long-Term Ecological Research Program in Puerto Rico, my lab is collaborating on a number of projects related to associations between taxonomic, functional, and phylogenetic biodiversity of soil microbes. We are interested in spatial variation in these associations along various gradients, including extensive elevational gradients that extends from sea level to ~1200 m asl. Because one transect passes through three forest types (tabonuco, palo colorado, and elfin forest) and the other parallel transect passes only through palm forest, we are partially able to decouple the effects of elevational changes in the flora from elevational changes in the abiotic environment. We are also documenting successional changes in the microbial biodiversity of communities in response to disturbances such as cyclonic storms and droughts. Finally, we are participating in a canopy trimming experiment that decouples confounded factors (detrital input and altered microclimate) associated with hurricane-induced gap formation on the biodiversity of microbial biodiversity.