Host-Microbe Interactions
Studying the interaction and co-evolution of beneficial and pathogenic microbes with host animals or plants.
Food Safety and Gut Health
Dr. Amalaradjou's research primarily focusses on food safety and gut health. Specifically, her research is targeted towards understanding pathogen survival and persistence along the food chain, their virulence attributes and the application of probiotics to control foodborne pathogens. She also studies the efficacy of currently applied intervention (hurdle) technologies to reduce pathogen transmission during pre- and post-harvest processing of fresh produce, meat and dairy products. Beyond controlling pathogens in food systems, her research also investigates the potential application of probiotics and functional foods in the prevention of gut pathologies. Major efforts in her lab are directed towards elucidating the molecular cross talk between diet, microbiome and host on all levels of regulation namely, genes, proteins and metabolites and ii) developing evidence-based dietary interventions for restoring health and preventing chronic pathologies. Through all aspects of her research, Dr. Amalaradjou's main objective is to improve food safety and promote public health.
Investigating survival strategies of bacterial persisters
Bacteria can utilize a diverse repertoire of strategies to survive stress, including those imposed by antimicrobials present in their natural habitats and in clinical settings. Research in the Mok Lab focuses on the survival strategies of bacterial persisters, which are rare subpopulations in clonal cultures that can survive antibiotic doses lethal to their kin. We use genetic, biochemical, and systems approaches to elucidate bacterial responses that contribute to antibiotic persistence and develop strategies to sabotage these culprits of recalcitrant infections.
Understanding drug metabolism by the gut microbiome
The Na Li lab is interested in studying how the human gut microbiome interact with orally ingested drug and bioactive compounds as well as their health outcomes, with an aim of improving the efficacy and minimizing interpersonal variations of oral drugs. We use a variety of chemical and biological approaches to study the role of the gut microbiome as an organ of metabolism.
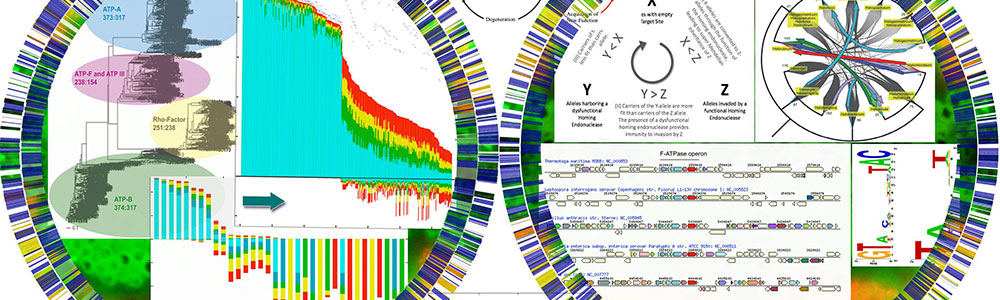
Research in Chris Simon's lab focuses on using information on molecular genetic processes to improve evolutionary tree-building (phylogenetics). My lab works at the interface between theory and observations to test newly proposed phylogenetic tree-building models and methods. We use phylogenetic trees to answer questions related to the origin, spread, maintenance, and documentation of biological diversity and how symbionts interact with hosts to facilitate or hinder colonization of new communities/adaptive zones.
Research in Daniel Bolnick's lab seeks to understand how ecological interactions affect the evolution of within-species trait variation. Research in the lab touches on a wide variety of species interactions, and combines theoretical models, natural history, field and lab experiments, and meta-analyses. Currently evolution of vertebrate immunity to parasites is a major, but not exclusive, focus of the lab.
Evan Ward’s laboratory studies the bivalve-associated microbiome to understand its involvement in host physiological processes. Bivalves are important members of coastal ecosystems and have commercial value, yet their microbiomes remain largely unexplored. We are interested in the role of gut/digestive gland and gill microbiota in bivalve toxicology, specifically involving anthropogenic pollutants like microplastics, nanoparticles, and pesticides. We utilize traditional bivalve husbandry and toxicological methods along with high-throughput sequencing and culture-based microbial techniques to conduct this research.
Benson's research interests include microbial biogeography, including the molecular and genomic interactions of bacteria with their environment, and the genomic evolution controlling distribution of microbes in several environments. The specific areas of interest include plant-microbe and insect-microbe microbiomes, and the distribution and diversity of microorganisms associated with cheese.
Early mortality and recurrent hospitalizations due to invasive pneumococcal disease (Streptococcus pneumoniae infection) are common for people with Sickle Cell Disease (SCD). Frequent pneumococcal vaccinations are administered to SCD patients in the US, reducing infections and increasing patient life expectancy. Unfortunately, recent research indicates that the currently licensed pneumococcal vaccines are not as effective in patients with SCD, which is associated with low antibody titers after vaccination. Our group has shown that mice with SCD are not protected from infection after pneumococcal vaccination. Much like in SCD patients, we correlated this lack of protection with short-lived pneumococcal antibody titers after vaccination. An important finding from this work is that antibody titers do not recover after booster shots are given to SCD mice, indicating that there is a deficiency in the generation of both B-cell memory and long-lived plasma cells (which produce antibodies). Further compounding this problem, it has been shown that protective pneumococcal vaccine responses require specialized B-cells in the spleen and peritoneal cavity called B-1 B-cells, and we have previously shown that SCD mice have dramatically reduced numbers of B-1 cells in their spleens. We also observed reduced percentages of these cells in the blood of SCD patients when compared to healthy control subjects. Hence, we have found cellular and functional correlates in the SCD mouse model that may explain the reduced effectiveness of pneumococcal vaccines and increased susceptibility to infection in people with SCD. We are currently working on understanding the basic mechanism of impaired pneumococcal immunity in SCD mice, with the aim of developing new vaccines and therapeutics to overcome this problem.
In collaboration with the Gage, Graf, Mustain, Noll, and Srivastava labs, Leslie Shor’s research group hasbeen developing systems to culture the microbiome of the lower termite Reticulitermes flavipes In the termite paunch, micro-scale oxygen gradients help to support a spatially-structured community capable of efficiently converting a mixed lignin-rich feedstock into usable chemicals such as acetate. Our approach is to construct micro-bioreactors with dynamically-controlled oxygen gradients using micro-manufacturing, electrolysis, and laminar flow patterning.
Dr. Hird is interested in the microbial biodiversity associated with vertebrates. Her laboratory predominantly studies microbiome samples from wild birds (although they are also interested in other taxa). We want to understand what microbes live with what birds, including when, where and why they do so. We use high-throughput amplicon sequencing, metagenomic and metatranscriptomic sequencing, and whole genome sequencing to document microbial partners; we use phylogenetic comparative methods to interpret results through an evolutionary lens.
Sarah Knutie is a disease ecologist and evolutionary biologist who studies the role of gut microbiota in host defenses against parasites. Specifically, she’s interested in whether and how the gut microbiota affects developmental immunity to micro- and macroparasites in birds, frogs, and fish. Notably, her lab works in the Galapagos Islands where she studies the effect of an emerging disease on Darwin’s finches and how urbanization can influence host defense mechanisms against the parasite.
Arthur Günzl’s research team focuses on gene expression mechanisms in the eukaryotic, unicellular parasite Trypanosoma brucei which differ substantially from the correponding mammalian processes, depend on extremely divergent protein factors, and enable the parasite to evade the mammalian immune system by Antigenic Variation. We use biochemical, genetic and imaging techniques to study these processes and factors. A particular strenght of our group is the purification of protein complexes by tandem affinity purification.
The Srivastava Group is interested in applying quantitative systems biology to understand gene regulatory networks, metabolic networks, and signaling networks. Machine learning algorithms in conjunction with reaction kinetics theory are used extensively to mine experimental data and various publically available databases (e.g. GenBank, PubMed, etc.) to elucidate network architecture, discover new pathways/interactions, and generate novel hypotheses. Potential applications include more robust treatment of infectious diseases and cancer, as well as advancing and optimizing industrial biotechnology processes.
Marcy Balunas’ lab investigates the chemistry and biological activity of marine microorganisms with a unique focus on the microbiology and natural products chemistry of tunicate-associated and psychrophilic marine and glacial bacteria. We are interested in the microbial dynamics in marine tunicates, especially as related to the sequestration, inheritance, and function of the bacterial communities associated with marine tunicates. Our approach includes both chemical (e.g., extraction, compound isolation, HPLC, LC-MS, NMR) and biological (e.g., pathogenic biological assays, 16S rRNA deep sequencing) techniques.
The Gage lab uses a combination of traditional genetics, molecular biology, biochemistry microscopy, and genome-scale technologies to study the interactions of microbes with plants and various kinds of protists. In addition, we develop tools for monitoring complex environments at small spatial scales. Areas of particular interest are: 1) cell-cell communication, 2) signal transduction, 3) the assembly and function microbial communities that contain both prokaryotes and protists and 4) developing biosensor technologies.
Steven Geary’s research interests are microbe-host interactions and evolution of microbial virulence, specifically involving bacterial pathogens of the genus Mycoplasma and primarily Mycoplasma gallisepticum; both in its natural poultry host and in the novel host, the American house finch. The lab uses comparative and functional genomics, transcriptomics, and in vivo models to investigate mechanisms of mycoplasmal pathogenesis, vaccine design, and virulence evolution. Website: http://patho.uconn.edu/people/faculty/indeste.html
The Nyholm laboratory explores beneficial host/microbe interactions, ultimately trying to understand the mechanisms by which bacterial symbionts and their animal hosts communicate and how the host’s innate immune system might influence these interactions. We are currently studying how these complex associations are established, regulated and maintained. We use the model association between the Hawaiian bobtail squid, Euprymna scolopes, and the bioluminescent bacterium Vibrio fischeri, to to understand mechanisms of host/symbiont specificity. We are also exploring a bacterial consortium that is found in a reproductive gland of female squid and is hypothesized to provide antimicrobial and/or antifouling compounds in order to protect the developing, externally laid embryos.
Jonathan Klassen studies microbial community ecology, especially using the fungus-growing ant symbiosis as a model system to understand how microbial interaction networks evolve. He is particularly interested in multipartite symbioses, which are common in nature and include complicated context-dependent dynamics that are not well described by more typical pair-wise symbiotic frameworks. The Klassen lab uses genomic, phenotypic and chemical biology techniques to describe the diversity, ecology and evolution of model multipartite symbioses, especially to identify keystone members and emergent properties of these networks that underpin their stability and function.
The Wunder lab work is focused on better understanding how the pathogenic Leptospira bacteria causes infection in susceptible and reservoir animals. Using animal models, in vitro assays, transcriptomics, comparative and functional genomics, and molecular biology techniques our goal is to identify targets of interest, understand their role on pathogenesis, and determine their mechanism of function. Ultimately, this knowledge is applied to identify and evaluate the potential of novel virulence factors as candidates for diagnostic and prevention of leptospirosis, an important neglected zoonotic disease, focusing on human and animal health.